[1]:
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
mpl.rcParams.update(
{
"text.usetex": False,
"axes.labelsize": 20,
"figure.labelsize": 18,
"xtick.labelsize": 16,
"ytick.labelsize": 16,
"figure.constrained_layout.wspace": 0,
"figure.constrained_layout.hspace": 0,
"figure.constrained_layout.h_pad": 0,
"figure.constrained_layout.w_pad": 0,
"axes.linewidth": 1.3,
}
)
import jax
import jax.numpy as jnp
# you should always set this
jax.config.update("jax_enable_x64", True)
Damped Harmonic Oscillator (DHO) Fitting
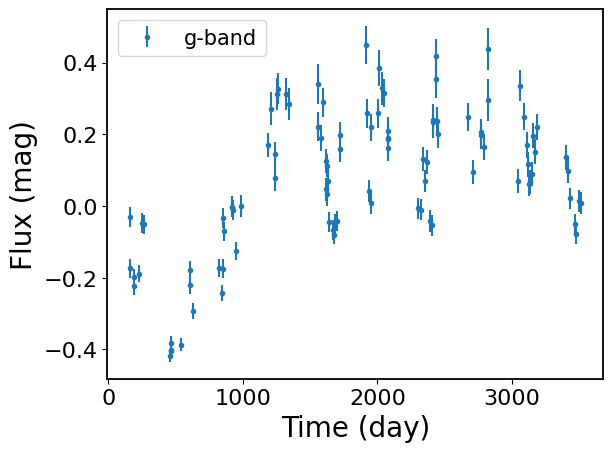
This notebook demonstrate how to fit a DHO process to a single-band light curve. Please see notebook 01_MultibandFitting for a tutorial on fitting multiband light curves.
Note: The CARMA autoregressive parameter notation follows the covention of Kelly+14.
1. Light Curve Simulation
[2]:
from eztao.carma import DHO_term
from eztao.ts import addNoise, gpSimRand
[3]:
# CARMA(2,1)/DHO parameters
alphas = {"g": [0.05, 0.0002]}
betas = {"g": [0.0006, 0.03]}
# simulation configureations
snrs = {"g": 5}
sampling_seeds = {"g": 2}
noise_seeds = {"g": 11}
lags = {"g": 0}
bands = "g"
n_yr = 10
ts, ys, yerrs = {}, {}, {}
ys_noisy = {}
seed = 2
for band in bands:
DHO_kernel = DHO_term(*np.log(alphas[band]), *np.log(betas[band]))
t, y, yerr = gpSimRand(
DHO_kernel,
snrs[band],
365 * n_yr, # 10 year LC
100,
lc_seed=seed,
downsample_seed=sampling_seeds[band],
)
# add to dict
ts[band] = t
ys[band] = y
yerrs[band] = yerr
# add simulated photometric noise
ys_noisy[band] = addNoise(ys[band], yerrs[band], seed=noise_seeds[band] + seed)
for b in bands:
plt.errorbar(
ts[b][::1], ys_noisy[b][::1], yerrs[b][::1], fmt=".", label=f"{b}-band"
)
plt.xlabel("Time (day)")
plt.ylabel("Flux (mag)")
plt.legend(fontsize=15)
[3]:
<matplotlib.legend.Legend at 0x7295bcccf410>
2. Fitting
Here, we demonstrate how to use the UniVarModel for fitting single-band light curves.
[4]:
import numpyro
import numpyro.distributions as dist
from eztaox.fitter import random_search
from eztaox.kernels.quasisep import CARMA
from eztaox.models import UniVarModel
from numpyro.handlers import seed as numpyro_seed
2.1 Initialize Light Curve Model
[5]:
zero_mean = False
p = 2 # CARMA p-order
test_params = {"log_kernel_param": jnp.log(np.array([0.1, 1.1, 1.0, 3.0]))}
# define kernel
k = CARMA.init(
jnp.exp(test_params["log_kernel_param"][:p]),
jnp.exp(test_params["log_kernel_param"][p:]),
)
# define univar model
m = UniVarModel(ts["g"], ys_noisy["g"], yerrs["g"], k, zero_mean=zero_mean)
m
[5]:
UniVarModel(
X=(f64[100], i64[100]),
y=f64[100],
diag=f64[100],
base_kernel_def=<jax._src.util.HashablePartial object at 0x7295bbf37d10>,
multiband_kernel=<class 'eztaox.kernels.quasisep.MultibandLowRank'>,
nBand=1,
mean_func=None,
amp_scale_func=None,
lag_func=None,
zero_mean=False,
has_jitter=False,
has_lag=False
)
2.2 Define InitSampler
[6]:
def initSampler():
# DHO Alpha & Beta parameters
log_alpha = numpyro.sample(
"log_alpha", dist.Uniform(low=-16.0, high=0.0).expand([2])
)
log_beta = numpyro.sample("log_beta", dist.Uniform(low=-10.0, high=2.0).expand([2]))
log_kernel_param = jnp.hstack([log_alpha, log_beta])
numpyro.deterministic("log_kernel_param", log_kernel_param)
# mean
mean = numpyro.sample("mean", dist.Uniform(low=-0.2, high=0.2))
sample_params = {"log_kernel_param": log_kernel_param, "mean": mean}
return sample_params
[7]:
# generate a random initial guess
sample_key = jax.random.PRNGKey(1)
prior_sample = numpyro_seed(initSampler, rng_seed=sample_key)()
prior_sample
[7]:
{'log_kernel_param': Array([ -4.87392318, -13.69996053, -3.87823238, -6.84304299], dtype=float64),
'mean': Array(-0.06222013, dtype=float64)}
2.3 MLE Fitting
[8]:
%%time
model = m
sampler = initSampler
fit_key = jax.random.PRNGKey(1)
nSample = 10_000
nBest = 10 # it seems like this number needs to be high
bestP, ll = random_search(model, initSampler, fit_key, nSample, nBest)
bestP
CPU times: user 3.05 s, sys: 176 ms, total: 3.23 s
Wall time: 3.24 s
[8]:
{'log_kernel_param': Array([-9.18107731, -2.83734073, -7.38433175, -3.38352044], dtype=float64),
'mean': Array(0.04086568, dtype=float64)}
[9]:
# True DHO param
# Note that EzTao follows the CARMA notation from Moreno+19,
# and EzTaoX adopts the CARMA notation from Kelly+14.
# The main difference is that the alpha parameter index is reversed.
print("True DHO Params (in natual log):")
print(np.log(np.hstack([alphas["g"][::-1], betas["g"]])))
print("MLE DHO Params (in natual log):")
print(bestP["log_kernel_param"])
True DHO Params (in natual log):
[-8.51719319 -2.99573227 -7.4185809 -3.5065579 ]
MLE DHO Params (in natual log):
[-9.18107731 -2.83734073 -7.38433175 -3.38352044]
3. MCMC
[10]:
import arviz as az
from numpyro.infer import MCMC, NUTS
[11]:
def numpyro_model(t, yerr, y=None):
log_alpha = numpyro.sample(
"log_alpha", dist.Uniform(low=-16.0, high=0.0).expand([2])
)
log_beta = numpyro.sample("log_beta", dist.Uniform(low=-10.0, high=2.0).expand([2]))
log_kernel_param = jnp.hstack([log_alpha, log_beta])
numpyro.deterministic("log_kernel_param", log_kernel_param)
# mean: use a normal prior for better convergence
mean = numpyro.sample("mean", dist.Normal(0.0, 0.1))
sample_params = {"log_kernel_param": log_kernel_param, "mean": mean}
# the following is different from the initSampler
zero_mean = False
p = 2
k = CARMA.init(
jnp.exp(test_params["log_kernel_param"][:p]),
jnp.exp(test_params["log_kernel_param"][p:]),
)
m = UniVarModel(ts["g"], ys_noisy["g"], yerrs["g"], k, zero_mean=zero_mean)
m.sample(sample_params)
[12]:
%%time
nuts_kernel = NUTS(
numpyro_model,
dense_mass=True,
target_accept_prob=0.9,
init_strategy=numpyro.infer.init_to_sample,
)
mcmc = MCMC(
nuts_kernel,
num_warmup=1000,
num_samples=5000,
num_chains=1,
# progress_bar=False,
)
mcmc_seed = 0
mcmc.run(jax.random.PRNGKey(mcmc_seed), ts["g"], yerrs["g"], y=ys_noisy["g"])
data = az.from_numpyro(mcmc)
mcmc.print_summary()
sample: 100%|██████████| 6000/6000 [00:13<00:00, 457.27it/s, 31 steps of size 9.03e-02. acc. prob=0.99]
mean std median 5.0% 95.0% n_eff r_hat
log_alpha[0] -10.05 2.11 -9.96 -13.45 -6.12 323.11 1.02
log_alpha[1] -2.70 0.93 -2.84 -4.30 -1.14 232.95 1.01
log_beta[0] -7.26 1.25 -7.42 -9.50 -5.20 232.07 1.02
log_beta[1] -3.38 0.49 -3.37 -3.71 -2.98 390.68 1.00
mean 0.02 0.08 0.02 -0.12 0.15 2287.80 1.00
Number of divergences: 0
CPU times: user 17.1 s, sys: 423 ms, total: 17.5 s
Wall time: 17.6 s
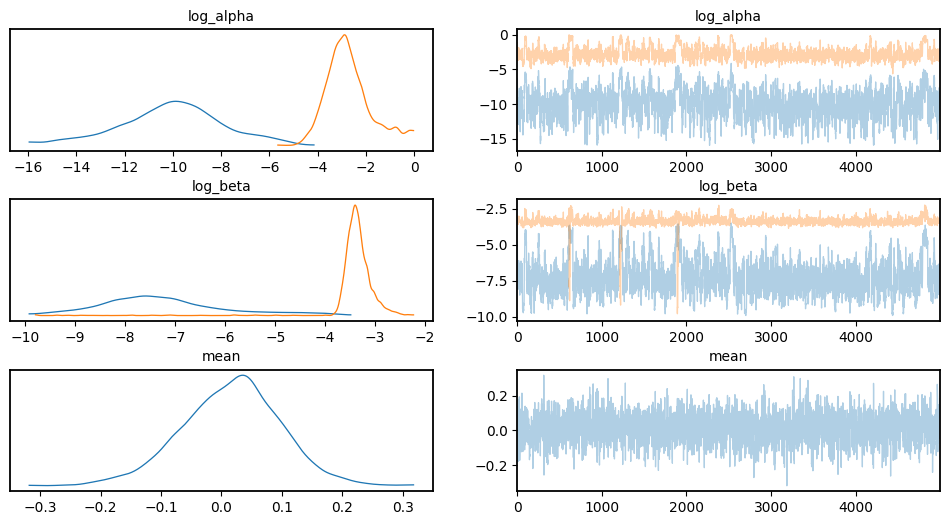
Visualize Chains, Posterior Distributions
[13]:
import warnings
warnings.filterwarnings("ignore", category=RuntimeWarning)
[14]:
az.plot_trace(data, var_names=["log_alpha", "log_beta", "mean"])
plt.subplots_adjust(hspace=0.4)
[15]:
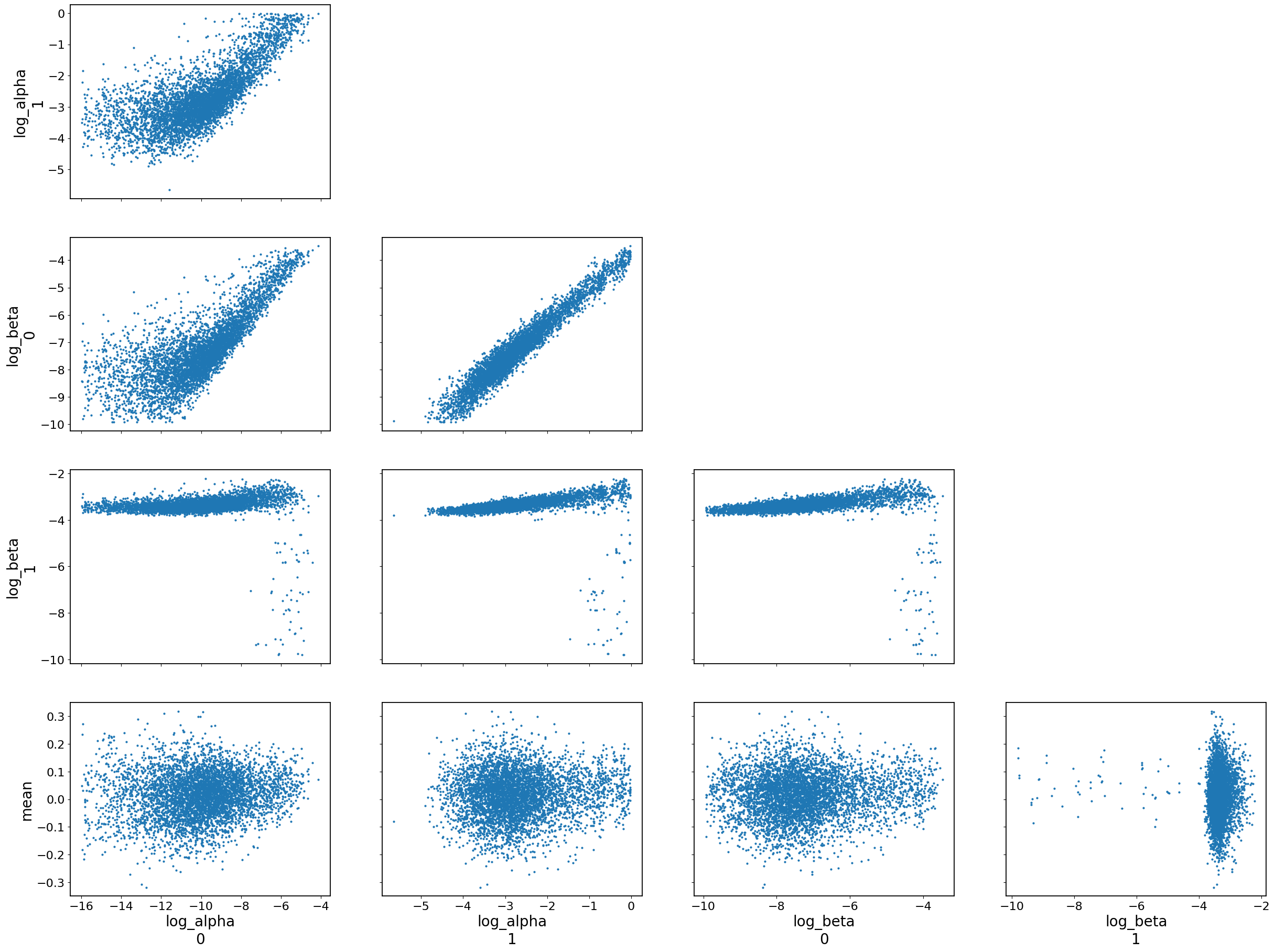
az.plot_pair(data, var_names=["log_alpha", "log_beta", "mean"])
[15]:
array([[<Axes: ylabel='log_alpha\n1'>, <Axes: >, <Axes: >, <Axes: >],
[<Axes: ylabel='log_beta\n0'>, <Axes: >, <Axes: >, <Axes: >],
[<Axes: ylabel='log_beta\n1'>, <Axes: >, <Axes: >, <Axes: >],
[<Axes: xlabel='log_alpha\n0', ylabel='mean'>,
<Axes: xlabel='log_alpha\n1'>, <Axes: xlabel='log_beta\n0'>,
<Axes: xlabel='log_beta\n1'>]], dtype=object)
4. Second-order Statistics
[16]:
from eztaox.kernel_stat2 import gpStat2
ts = np.logspace(0, 4)
fs = np.logspace(-4, 0)
[17]:
# get MCMC samples
flatPost = data.posterior.stack(sample=["chain", "draw"])
log_carma_draws = flatPost["log_kernel_param"].values.T
[18]:
# create second-order stat object
dho_k = CARMA.init(alphas["g"][::-1], betas["g"])
gpStat2_dho = gpStat2(dho_k)
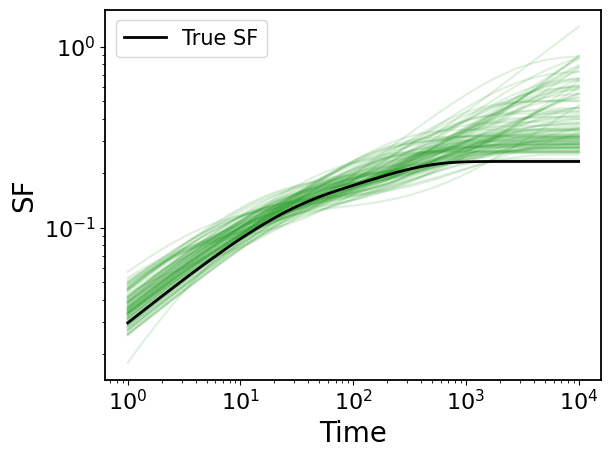
4.1 Structure Function
[19]:
# compute sf for MCMC draws
mcmc_sf = jax.vmap(gpStat2_dho.sf, in_axes=(None, 0))(ts, jnp.exp(log_carma_draws))
[20]:
## plot
# ture SF
plt.loglog(ts, gpStat2_dho.sf(ts), c="k", label="True SF", zorder=100, lw=2)
plt.legend(fontsize=15)
# MCMC SFs
for sf in mcmc_sf[::50]:
plt.loglog(ts, sf, c="tab:green", alpha=0.15)
plt.xlabel("Time")
plt.ylabel("SF")
[20]:
Text(0, 0.5, 'SF')
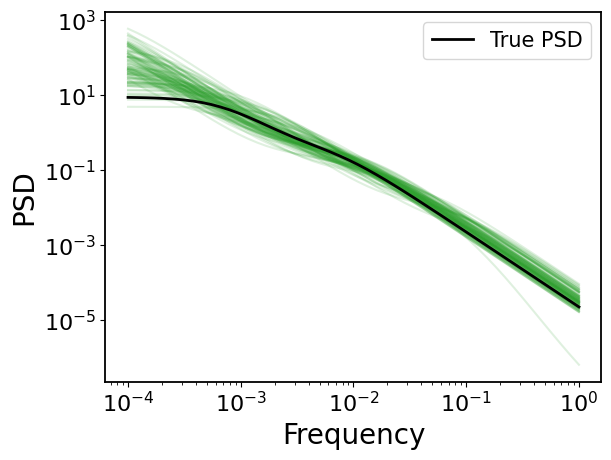
4.1 Power Spectral Density (PSD)
[21]:
# compute sf for MCMC draws
mcmc_psd = jax.vmap(gpStat2_dho.psd, in_axes=(None, 0))(fs, jnp.exp(log_carma_draws))
[22]:
## plot
# ture PSD
plt.loglog(fs, gpStat2_dho.psd(fs), c="k", label="True PSD", zorder=100, lw=2)
plt.legend(fontsize=15)
# MCMC PSDs
for psd in mcmc_psd[::50]:
plt.loglog(fs, psd, c="tab:green", alpha=0.15)
plt.xlabel("Frequency")
plt.ylabel("PSD")
[22]:
Text(0, 0.5, 'PSD')
[ ]: